Personal Website
Research with undergrads at VCU
Looking for a reseach opportunity in bioinformatics?

You are in the correct place!
Email GB with your interest to get started:
gbbulut at vcu.edu
Research with undergrads at William & Mary
GB's Postdoctoral Research
Illuminating, profiling and manipulating perivascular plasticity in microvasculature
-
My postdoc project focused on adipose tissue microvasculature and the role of smooth muscle cell (SMC)- derived cells in the context of diet induced obesity. I performed extensive flow cytometric and scRNA sequencing based investigation of cell types in adipose tissue microvasculature. (You can read more here.)
-
Smooth Muscle Cells (SMC) make up the involuntary muscles in our body and cover many hollow organs and the vascular system such as arteries, veins, and lymph vessels. Cardiovascular diseases are the leading cause of death due to heart attacks and strokes. Chronic inflammation developing in the adipose tissue of an overweight or obese body also increase cardiovascular risks.
-
Adipose tissue microvasculature consists of highly heterogenous types of stromovascular cell types including macrophages and vascular cells. Interestingly, some macrophage types were increased in response to obesity, and decreased in response to smooth muscle specific knock out of pluripotency factor Klf4.
-
Interestingly, lymphatic smooth muscle cells were increased upon obesity along with decreases in blood pressure regulating SMC clusters.
-
Understanding cellular heterogeneity in adipose tissue and how it changes in response to obesity is of significant biomedical impact and could lead to novel therapies for diabetes and obesity related microvascular complications.
Macrophage
Heterogeneity


Lymphatic
smooth muscle cells

Obesity promoting
EndoMT

Human blood monocyte derived macrophages
Human blood monocytes can be utilized to differentiate and generate macrophage (hBMDM) in tissue culture. This system is highly physiologically relevant and a valuable platform to test molecular, cellular mechanisms related to macrophage biology.
I aim to use cultured monocyte derived macrophages to recapitulate the differentiated ATM subsets.
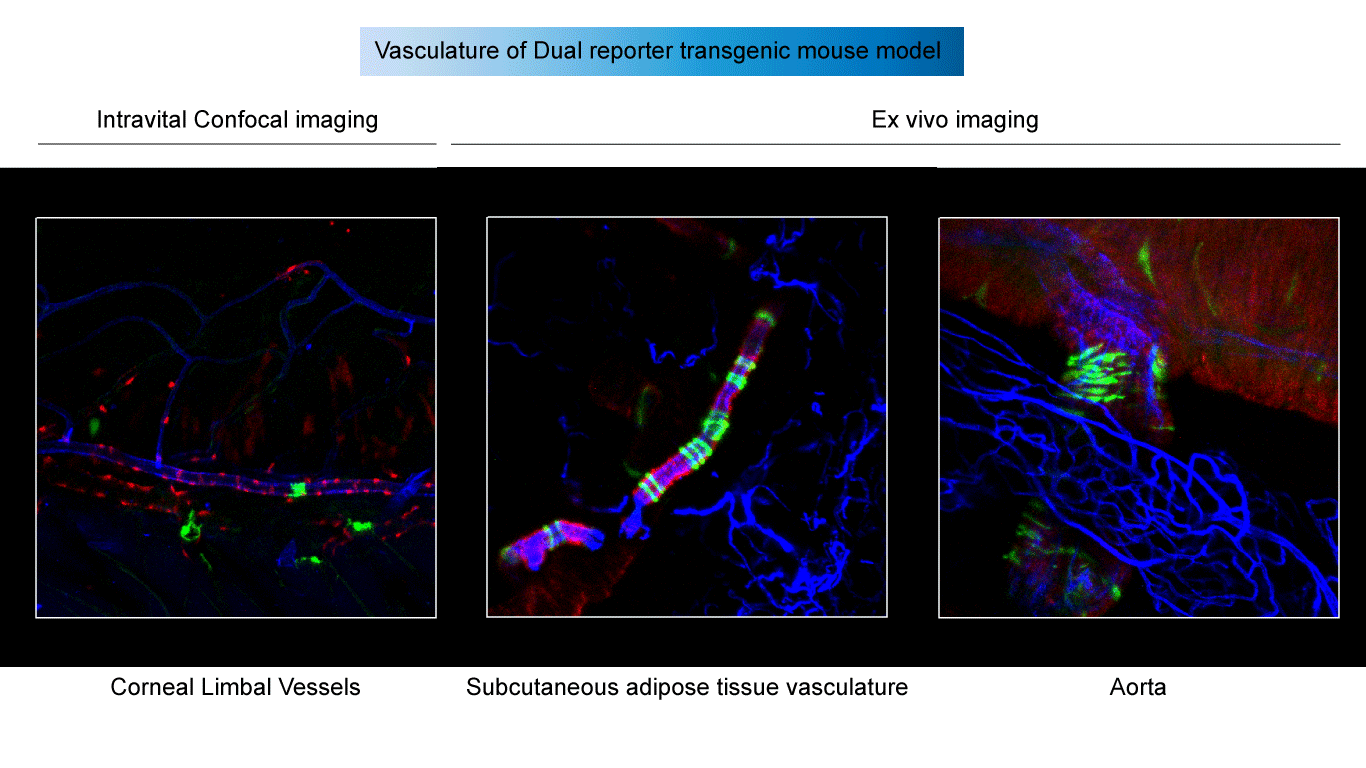
I have used Myh11-Lgals3 dual lineage tracing system for intravital and ex vivo imaging to illuminate SMC plasticity.
In the future, it would be a unique research avenue to create a sequential lineage tracer mouse model using Myh11 and Tnnt1-2 to label lymphatic smooth muscle cells which can be used for making knockouts for metabolic experiments and live imaging.
Overall, it could help us to know how we can preserve the health and well-being of macrophages, endothelial cells and vasculature as the adipose tissue transforms during obesity, to prevent dysregulation of blood pressure and vascular permeability.
Previously mentored students
Kritihika Layagala, undergrad at WM
Lillian Waller, Medical Resident at VCU
Ana Tsiskarishvili, District Manager at Vector Marketing
Sophia Kirmani, UVA graduate.











